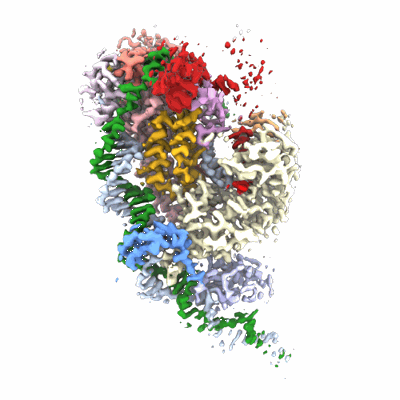
Cryo-EM structure of MLE in complex with SL7UUC RNA and ADP
Resolution: 4.24 Å
EM Method: Single-particle
Fitted PDBs: 8pjj
Q-score: 0.218
To Be Published
- Complex of mle, sl7uuc and adp (150 kDa, Complex)
- Sl7uuc rna (Complex from Drosophila melanogaster)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Dosage compensation regulator (Complex from Drosophila melanogaster)
- Sl7uuc rna (19 kDa, RNA from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
Cryo-EM structure of MLE in complex with UUC RNA and ADP
Resolution: 3.62 Å
EM Method: Single-particle
Fitted PDBs: 8pjb
Q-score: 0.415
To Be Published
- Rna (5'-r(p*cp*up*cp*up*up*up*cp*up*u)-3') (3 kDa, RNA from Drosophila melanogaster)
- Magnesium ion (24 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Complex of mle, uuc and adp (134 kDa, Complex)
- Dosage compensation regulator (Complex from Drosophila melanogaster)
- Rna (5'-r(p*cp*up*cp*up*up*up*cp*up*u)-3') (Complex from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
Cryo-EM structure of MLE in complex with ADP:AlF4 and U10 RNA
Resolution: 2.86 Å
EM Method: Single-particle
Fitted PDBs: 8b9g
Q-score: 0.513
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ Pubmed: 37989319 DOI: doi:10.1016/j.molcel.2023.10.026 ]
- Mle in complex with adp:alf4 and u10 rna (133 kDa, Complex from Drosophila melanogaster)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Rna (5'-r(p*up*up*up*up*up*up*up*up*up*up*u)-3') (3 kDa, RNA from Drosophila melanogaster)
- Water (18 Da, Ligand)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
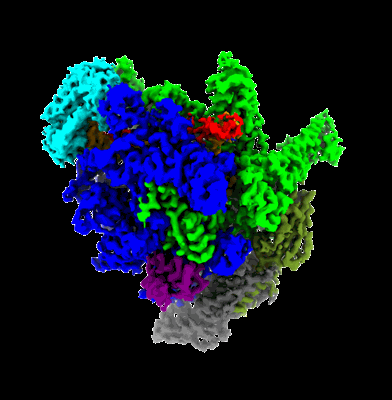
Cryo-EM structure of MLE in complex with ADP:AlF4 and UUC RNA
Resolution: 2.95 Å
EM Method: Single-particle
Fitted PDBs: 8b9i
Q-score: 0.469
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ Pubmed: 37989319 DOI: doi:10.1016/j.molcel.2023.10.026 ]
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Mle in complex with adp:alf4 and uuc rna (133 kDa, Complex from Drosophila melanogaster)
- Magnesium ion (24 Da, Ligand)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Water (18 Da, Ligand)
- Rna (5'-r(p*cp*cp*up*cp*up*up*up*cp*up*up*u)-3') (3 kDa, RNA from Drosophila melanogaster)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
Cryo-EM structure of MLE in complex with ADP:AlF4 and SL7modUUC RNA
Resolution: 4.04 Å
EM Method: Single-particle
Fitted PDBs: 8b9k
Q-score: 0.239
Jagtap PKA, Muller M, Kiss AE, Thomae AW, Lapouge K, Beck M, Becker PB, Hennig J
Mol Cell (2023) 83 pp. 4318-4333.e10 [ Pubmed: 37989319 DOI: doi:10.1016/j.molcel.2023.10.026 ]
- Mle in complex with adp:alf4 and sl7moduuc rna (149 kDa, Complex from Drosophila melanogaster)
- Tetrafluoroaluminate ion (102 Da, Ligand)
- Sl7moduuc (19 kDa, RNA from Drosophila melanogaster)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Dosage compensation regulator (130 kDa, Protein from Drosophila melanogaster)
Resolution: 3.1 Å
EM Method: Single-particle
Fitted PDBs: 7uwh
Q-score: 0.458
Hao Z, Gowder M, Proshkin S, Bharati BK, Epshtein V, Svetlov V, Shamovsky I, Nudler E
Cell (2023) 186 pp. 2425- [ DOI: doi:10.1016/j.cell.2023.04.029 Pubmed: 37196657 ]
- Dna-directed rna polymerase subunit beta' (155 kDa, Protein from Escherichia coli)
- Rna (18-mer) (5 kDa, RNA from Escherichia coli)
- Dna-directed rna polymerase subunit omega (10 kDa, Protein from Escherichia coli)
- Zinc ion (65 Da, Ligand)
- Dna-directed rna polymerase subunit beta (150 kDa, Protein from Escherichia coli)
- Elongation complex(ec)-rnaseh2 complex bound to ribonucleotide substrate (Complex from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- Ribonuclease hii (21 kDa, Protein from Escherichia coli)
- Dna-directed rna polymerase subunit alpha (36 kDa, Protein from Escherichia coli)
- Dna (59-mer) (18 kDa, DNA from Escherichia coli)
- Dna/rna (59-mer) (18 kDa, Macromolecule from Escherichia coli)
DNA initiation subcomplex of Xenopus laevis DNA polymerase alpha-primase
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8g9l
Q-score: 0.453
Mullins EA, Salay LE, Durie CL, Bradley NP, Jackman JE, Ohi MD, Chazin WJ, Eichman BF
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01227-4 Pubmed: 38491139 ]
- 2'-deoxyguanosine-5'-triphosphate (507 Da, Ligand)
- Primase (Complex from Xenopus laevis)
- Polymerase alpha (Complex from Xenopus laevis)
- Polymerase alpha-primase with a dna initiation substrate (110 kDa, Complex)
- Dna primase large subunit (59 kDa, Protein from Xenopus laevis)
- Rna primer (2 kDa, RNA from synthetic construct)
- Iron/sulfur cluster (351 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Dna polymerase alpha catalytic subunit (129 kDa, Protein from Xenopus laevis)
- Dna initiation substrate (20 kDa, Complex from synthetic construct)
- Dna template (15 kDa, DNA from synthetic construct)
In situ structure of polymerase complex of mammalian reovirus in the elongation state
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 7yed
Q-score: 0.498
Bao K, Zhang X, Li D, Sun W, Sun Z, Wang J, Zhu P
PNAS (2022) 119 pp. e2203054119-e2203054119 [ DOI: doi:10.1073/pnas.2203054119 Pubmed: 36469786 ]
- Rna (Complex)
- Mu-2 protein (83 kDa, Protein from Mammalian orthoreovirus 3)
- Lambda-2 protein (143 kDa, Protein from Mammalian orthoreovirus 3)
- Rna (5'-r(p*up*up*up*ap*ap*ap*ap*ap*up*up*up*up*ap*ap*ap*ap*up*ap*ap*up*u)-3') (6 kDa, RNA from Mammalian orthoreovirus 3)
- Rna (36-mer) (11 kDa, RNA from Mammalian orthoreovirus 3)
- Zinc ion (65 Da, Ligand)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Rna helicase (141 kDa, Protein from Mammalian orthoreovirus 3)
- Uridine 5'-triphosphate (484 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- S-adenosylmethionine (398 Da, Ligand)
- Mammalian orthoreovirus 3 (Virus from Mammalian orthoreovirus 3)
- Mammalian orthoreovirus 3 (Complex)
- Rna-directed rna polymerase (142 kDa, Protein from Mammalian orthoreovirus 3)
Bombyx mori R2 retrotransposon initiating target-primed reverse transcription
Resolution: 3.08 Å
EM Method: Single-particle
Fitted PDBs: 8gh6
Q-score: 0.49
Wilkinson ME, Frangieh CJ, Macrae RK, Zhang F
Science (2023) 380 pp. 301-308 [ DOI: doi:10.1126/science.adg7883 Pubmed: 37023171 ]
- Thymidine-5'-triphosphate (482 Da, Ligand)
- 28s dna bottom strand, 5' side (priming strand) (7 kDa, DNA from Bombyx mori)
- Zinc ion (65 Da, Ligand)
- 28s dna bottom strand, 3' side (21 kDa, DNA from Bombyx mori)
- R2bm 3'utr rna (81 kDa, RNA from Bombyx mori)
- 28s dna top strand (21 kDa, DNA from Bombyx mori)
- Magnesium ion (24 Da, Ligand)
- Bombyx mori r2 retrotransposon initiating target-primed reverse transcription (230 kDa, Complex from Bombyx mori)
- Reverse transcriptase-like protein (123 kDa, Protein from Bombyx mori)
Porcine epidemic diarrhea virus core polymerase complex
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8g6r
Q-score: 0.435
Anderson TK, Hoferle PJ, Lee KW, Coon JJ, Kirchdoerfer RN
bioRxiv (2023) [ Pubmed: 36993498 DOI: doi:10.1101/2023.03.15.532841 ]
- Nsp7 (6 kDa, Protein from Porcine epidemic diarrhea virus)
- Rna (5'-r(p*ap*ap*gp*ap*ap*gp*cp*up*ap*up*up*ap*ap*ap*ap*up*cp*ap*cp*a)-3') (6 kDa, RNA from synthetic construct)
- Core replication complex of porcine epidemic diarrhea virus composed of nsp12, nsp7, nsp8, and a short rna substrate (185 kDa, Complex from Porcine epidemic diarrhea virus)
- Zinc ion (65 Da, Ligand)
- Nsp8 (12 kDa, Protein from Porcine epidemic diarrhea virus)
- Rna (5'-r(p*gp*gp*up*up*gp*up*gp*ap*up*up*up*up*ap*ap*up*ap*gp*cp*up*u)-3') (6 kDa, RNA from synthetic construct)
- Nsp12 (104 kDa, Protein from Porcine epidemic diarrhea virus)